The human adaptive immune system is a natural diagnostic and therapeutic. Disease diagnostic information and therapeutic potential is directly encoded into antibody and T-cell immune repertoires. The advent of of high-throughput sequencing has enabled an unprecedented accumulation of big immune repertoire sequencing data (immunegenotype). However, as of yet, we lack the computational and experimental methods that help us decode the immune grammar that translates immunegenotype to immune state diagnosis and prediction of antigen binding. We believe that learning to read and write the immune repertoire language is key for the development of entirely novel, nature-inspired precision medicine immunodiagnostics and immunotherapeutics.
OVERVIEW
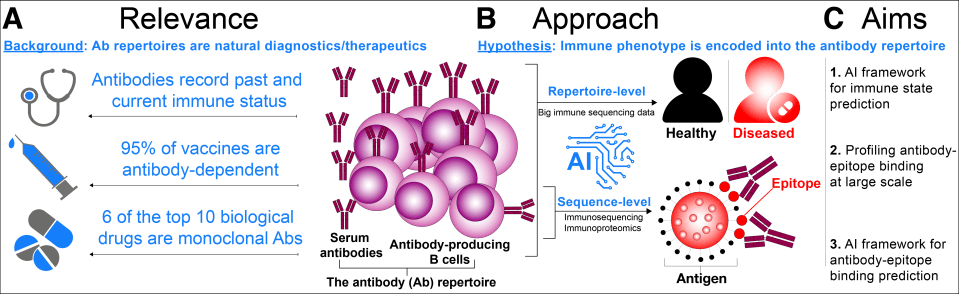
Our projects are grouped into three main areas of research and development:
- artificial intelligence methods for deciphering the human immune repertoire language
- new bioinformatics platforms for high-dimensional immune repertoire analysis,
- single-cell immune repertoire and immuno-mass spectrometry methods
We apply the developed methods to decipher the specific immune repertoire architecture of a wide array of human diseases spanning autoimmune disease (RA, T1D, Celiac disease), infection (Tuberculosis, Influenza) to cancer (GBM).
Our work is funded by the University of Oslo and UiO:LifeScience.
